Summary
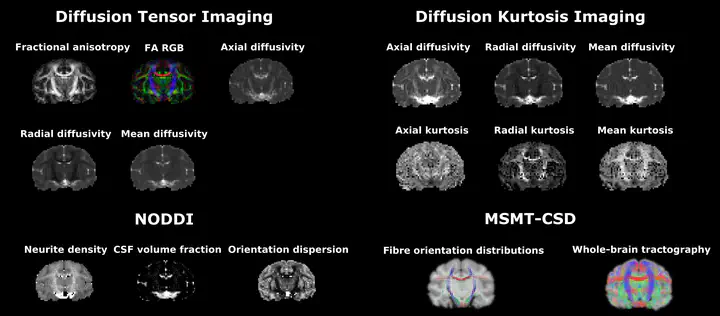
Diffusion MRI (dMRI) has the ability to link the MRI measurements at a millimeter-scale to tissue microstructures on a scale much smaller than the nominal image resolution. The reason for this is that the measured MR signal originates from the random motion of water molecules in the brain tissue at the cellular level. Various mathematical models have been proposed to decipher the tissue properties from the obtained dMRI signal. In this project, I provide python notebooks to calculate Diffusion Tensor Imaging (DTI), Diffusion Kurtosis Imaging (DKI), Neurite Orientation Dispersion and Density Imaging (NODDI), Single-Shell 3-tissue Constrained Spherical Deconvolution (SS3T-CSD), and Multi-Shell Multi-Tissue Constrained Spherical Deconvolution (MSMT-CSD) modeled parametric maps.